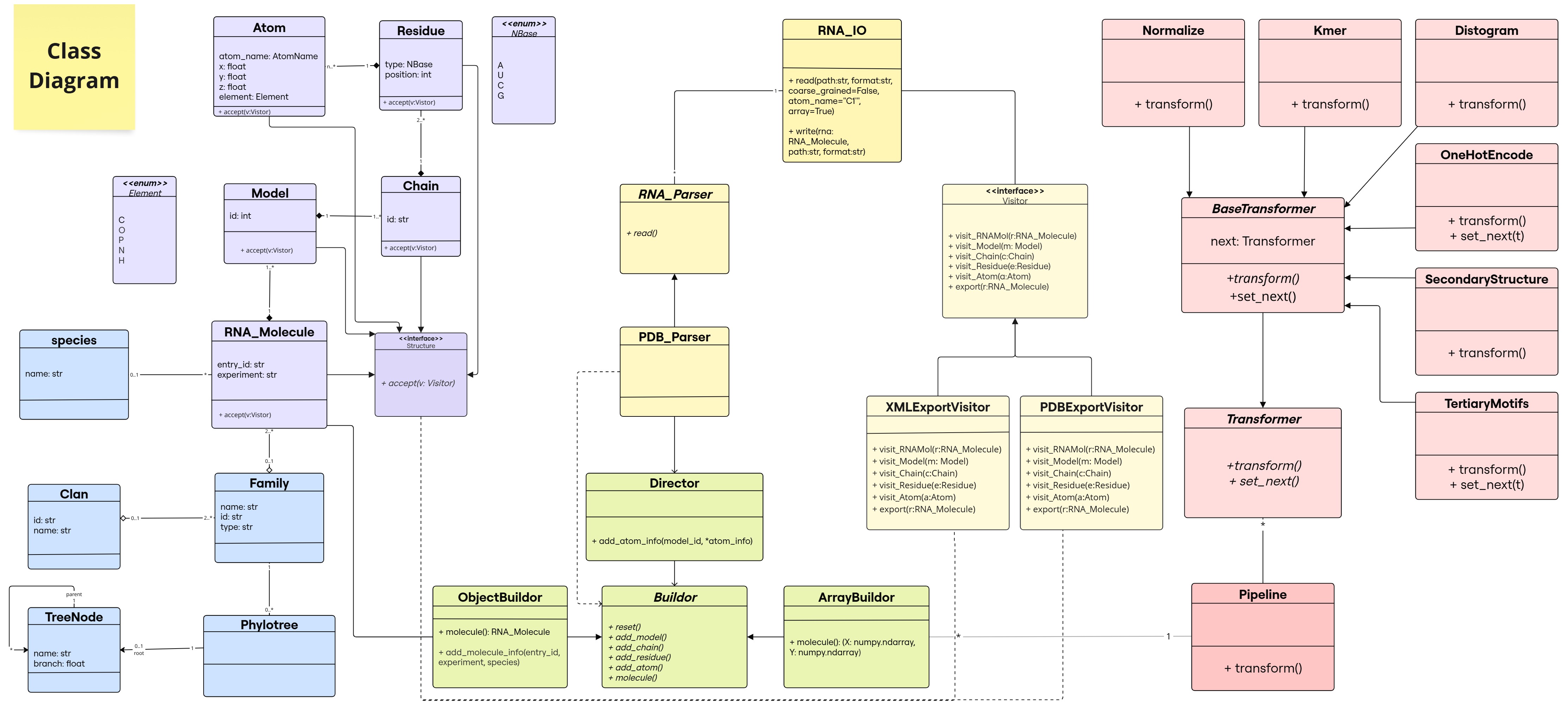
RNAr library
A python library allowing easy manipulation, reading, writing and transforming RNA structures to downstream analysis and training.
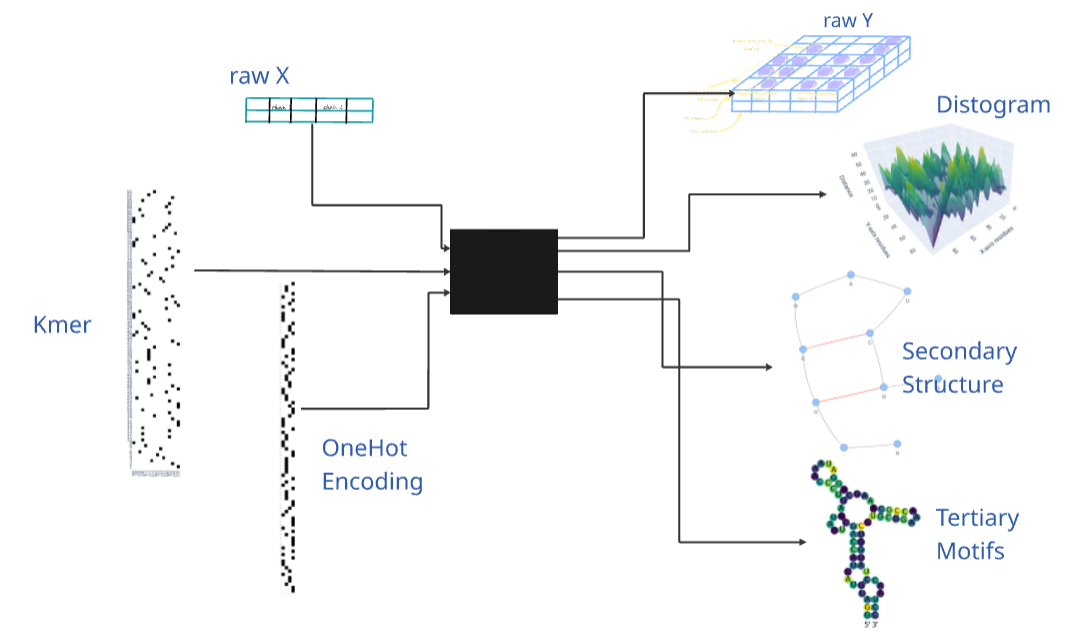
The latter image creates new horizon to view where this library is heading. Machine learning and deep learning are the new trends in bioinformatics, particulalry in structural prediction in the last couple of years, and this library is a step towards that direction. RNA structure prediction is still an open challenge today, many efforts are being made to present different types of features or labels to this model. The transformations in this library provide a good starting point to train models on different kinds of features and be assessed by the way the model is able to come near one of the RNA structural representation.
Installation
This library can be installed directly from github through pip:
pip install git+https://github.com/rna-oop/2425-m1-geniomhe-group-6.git
or simply clone the repository and install it locally:
git clone https://github.com/rna-oop/2425-m1geniomhe-group-6
cd 2425-m1geniomhe-group-6
pip install .
To make sure it’s installed correctly, you can run the following command in your terminal:
python -c "import RNAr; print(RNAr.__version__)"
Aknowledgements
| This project was developed as a sequence of labs from the Object-Oriented Programming (II) course given by Prof. Massinissa Hamidi at Université Evry Paris-Saclay , M1 bioinformatics | GENIOMHE-AI 2024/25. |
Group members:
Labs
Lab1
description | report | contents
Lab2
description | report | contents
Lab3
description | report | contents
Lab4
description | report | contents
Final Design